Why Cell-Based Protein Expression Isn't Always the Answer
If you've ever waited two weeks for a protein that never expressed, you already understand the fundamental limitations of traditional cell-based systems. Gene cloning, transformation, colony selection, culture optimization, and purification—each step introduces bottlenecks and potential failure points. For challenging targets like toxic proteins, membrane proteins, or proteins requiring non-standard amino acids, these limitations can make expression impossible.
Cell-free protein synthesis (CFPS) offers a fundamentally different approach that eliminates many of these constraints while dramatically accelerating timelines.
What is Cell-Free Protein Synthesis?
Cell-free protein synthesis, also called in vitro protein synthesis, leverages the cellular machinery responsible for translation—ribosomes, tRNAs, aminoacyl-tRNA synthetases, and translation factors—without requiring intact, living cells. Instead of transforming a plasmid into bacteria and waiting days for culture growth, CFPS systems combine purified components, or cell extracts with your DNA or mRNA template in a single reaction vessel.
The approach is straightforward: add your gene template to a mixture containing all the molecular machinery needed for transcription and translation. Protein synthesis begins immediately and typically completes within hours.
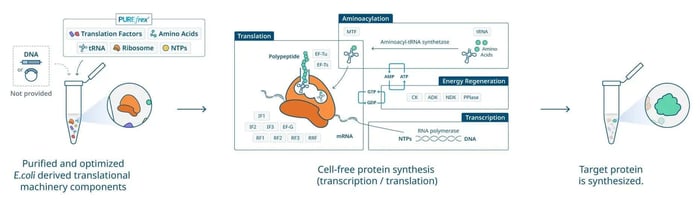 Figure 1. The in vitro, cell-free protein synthesis workflow.
Figure 1. The in vitro, cell-free protein synthesis workflow.Five Compelling Advantages of Cell-Free Protein Expression Systems
1. Dramatic Speed Advantage: Hours Instead of Days
Traditional expression workflows in mammalian cells or E. coli are inherently time-consuming, requiring gene cloning and vector construction, transformation and colony screening, culture growth and optimization, induction and harvest, followed by purification. Each step introduces delays, and the process typically requires 3-4 days or more from gene to purified protein (Figure 1). These extended timelines create bottlenecks in research projects, slow down screening campaigns, and delay critical experimental results.
Cell-free synthesis eliminates most of these steps. A DNA or mRNA template can be added directly to the reaction mixture, protein synthesis completes within 2-4 hours, and optional purification takes only a few additional hours. The entire workflow — from gene to protein — is completed in just 2-4 hours (Figure 2). This 10-fold reduction in timeline is particularly valuable for high-throughput screening, rapid prototyping, and time-sensitive research projects.
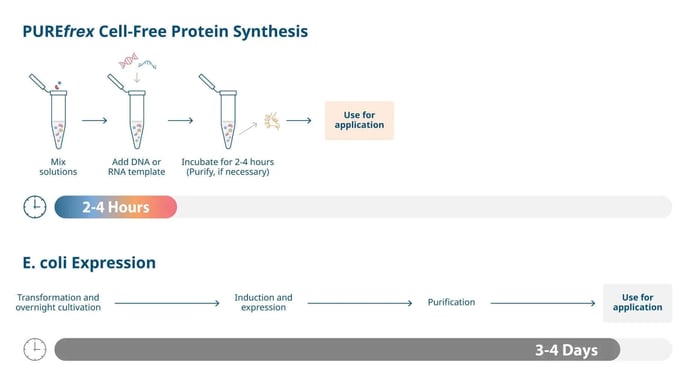 Figure 2. Comparison of workflow timelines, traditional E. coli expression vs PUREfrex® cell-free protein synthesis.
Figure 2. Comparison of workflow timelines, traditional E. coli expression vs PUREfrex® cell-free protein synthesis.2. Express Toxic Proteins Without Viability Concerns
Some of the most interesting proteins—antimicrobial peptides, proteases, membrane-disrupting proteins—are inherently toxic to host cells. Traditional expression fails because toxic proteins kill the host before sufficient yields can be achieved.
Cell-free systems eliminate this problem entirely. With no living cells to damage, toxicity becomes irrelevant. Proteins that are impossible to express in vivo can be produced consistently and at sufficient yields.
3. Complete Control Over Reaction Conditions
Unlike cell-based systems where your reaction occurs behind a cell membrane, CFPS reactions are completely open and accessible. You can:
- Monitor synthesis in real-time through sampling
- Adjust pH, salt concentration, or cofactor levels during the reaction
- Add specialized reagents like molecular chaperones, detergents for membrane proteins, or non-canonical amino acids
- Eliminate interference from cellular metabolism or competing pathways
This level of experimental control is impossible with intact cells and enables applications that cell-based systems simply cannot support.
4. Streamlined Workflow Enables Higher Throughput
Cell-free expression removes time-consuming steps: no cloning into expression vectors, no transformation, no colony screening, no culture optimization, no cell lysis. Simply prepare your template with the correct regulatory elements (T7 promoter, ribosome binding site, terminal stop codon) using standard PCR methodology, add it to the reaction mix, and incubate.
This simplified workflow is ideal for parallel screening of multiple constructs, rapid testing of mutations, and functional genomics studies requiring dozens or hundreds of proteins.
5. Unique Applications Traditional Methods Cannot Address
Membrane Protein Production
Add liposomes or nanodiscs directly to the cell-free reaction. Nascent membrane proteins insert co-translationally into lipid bilayers, maintaining proper topology and function. This approach successfully produces complex assemblies like SecYEG translocon and ATP synthase that typically misfold or aggregate in traditional systems.
Non-Standard Amino Acid Incorporation and Labeling
The open reaction environment allows direct addition of modified aminoacyl-tRNAs and unnatural amino acids. Specific protein labeling with fluorophores, biotin, radiolabels, or heavy atoms empowers structural biology studies, protein-protein interaction studies, and single-molecule analysis.
Disulfide Bond Formation
Supplement reactions with disulfide bond isomerases (DsbC) or eukaryotic oxidoreductase systems (PDI/Ero1α) to form native disulfide bonds—critical for antibody fragments, secretory proteins, and therapeutic candidates.
Key Research Applications
Cell-free protein synthesis excels in specific research scenarios:
Rapid Validation & Characterization
- Verify open reading frame (ORF) functionality before committing to large-scale expression
- Screen mutant libraries for expression and function
- Generate truncation variants for epitope mapping or domain analysis
- Produce proteins for interaction studies without weeks of optimization
Challenging Expression Targets
- Toxic proteins (antimicrobials, proteases, membrane-disrupting peptides)
- Membrane proteins (GPCRs, ion channels, transporters)
- Aggregation-prone proteins when supplemented with molecular chaperones
- Proteins with unusual amino acid compositions (poly-proline motifs, charged clusters)
Protein Engineering & Specialized Labeling
- Incorporate non-standard amino acids for biophysical probes
- Site-specific isotope labeling (¹⁵N, ¹³C, ²H) for NMR studies
- Fluorescent labeling via modified tRNAs for single-molecule studies
- Ribosome display for protein evolution studies
Synthetic Biology Applications
- Prototype synthetic circuits and biosensors
- Test gene network designs without cellular context
- Produce toxic or growth-inhibiting proteins for pathway engineering
- Generate components for cell-free metabolic systems
The PUREfrex® Advantage: Reconstructed Precision
While crude cell-free extracts have been available for decades, they suffer from significant limitations: contaminating nucleases and proteases, undefined component ratios, batch-to-batch variability, and background protein contamination. These issues compromise reproducibility and complicate downstream applications.
PUREfrex represents the next generation of cell-free technology—a fully reconstituted system using 36 individually purified components from E. coli translation machinery. Unlike traditional S30 extracts, every enzyme, ribosome, and translation factor in PUREfrex is defined, purified, and optimized.
Download our PUREfrex brochure
What Makes PUREfrex Different?
Exceptional Purity
Advanced purification protocols eliminate contaminating RNase, proteases, and β-galactosidase. Lipopolysaccharide (LPS) concentrations are reduced to ≤0.1 Endotoxin Units (EU) per microliter — critical for applications requiring high purity, such as therapeutic protein production or sensitive biological assays.
Superior Reproducibility
Defined component stoichiometry and quality control ensure consistent performance across batches. Unlike extract-based systems where each preparation varies, PUREfrex delivers predictable results.
Clean Background
No His-tags on system components means your protein purification isn't complicated by tagged translation factors. This design flexibility allows unrestricted template design—add any tag you need or produce completely untagged proteins.
Enhanced Performance
PUREfrex 2.0 and 2.1 deliver significantly higher yields than earlier versions: 600 μg/mL for DHFR and 800 μg/mL for GFP. These improvements translate directly to lower costs per experiment and higher throughput.
Flexible Enhancement with Targeted Supplements
PUREfrex's modular design allows targeted optimization for challenging proteins:
- Molecular Chaperones: DnaK Mix or GroE Mix improve folding and solubility of aggregation-prone targets.
- Redox Systems: DsbC Set or PDI Set enable proper disulfide bond formation in proteins that require them for stability and function.
- Translation Enhancers: EF-P Supplement facilitates translation through difficult sequences like poly-proline motifs.
This supplement strategy allows researchers to optimize reactions for specific protein properties without reformulating the entire system.
Moving Forward: Is PUREfrex Right for Your Research?
PUREfrex offers significant flexibility for precision protein engineering and scalable production while delivering reproducible, high-quality results. The combination of speed (10-fold faster), toxicity tolerance, experimental control, simplified workflows, and unique capabilities makes PUREfrex the logical choice for:
- Proteins toxic to host cells
- Rapid screening campaigns requiring high throughput
- Membrane proteins and other difficult targets
- Specialized labeling
- Research requiring quick turnaround of results
Ready to accelerate your protein expression workflow?
Learn more about the PUREfrex Cell-Free Protein Synthesis System and available supplements or place an order.
Interested in customizing PUREfrex for your specialized protein expression needs? We offer the development of custom PUREfrex kits that are adapted to your research goals. Request a technical consultation.